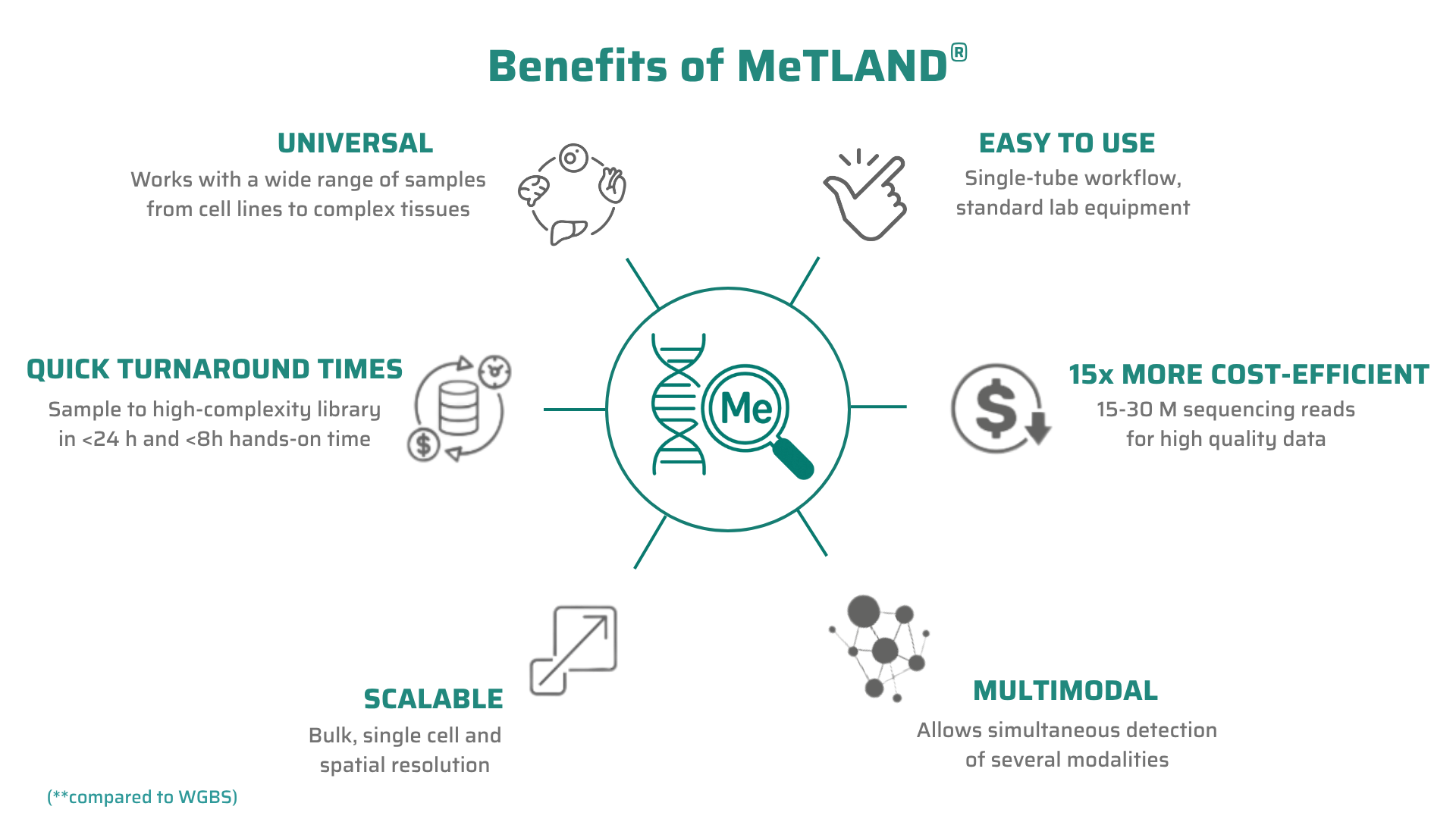
Our flagship product targets the deepest layer of the epigenome - DNA methylation. It uses our proprietary MeTLAND technology to capture methylation patterns across the genome in a way that is cheaper, more data-rich and easier to implement compared to existing alternatives.
Interested in MeTLAND®?
Try our Early Access Program
Our early access partners can:
Benchmark MeTLAND side-by-side with their current method
Test challenging or low-input samples
Generate high-complexity methylation libraries with fast turnaround
Provide feedback that shapes the first commercial release
This program is designed for teams who need reliable, scalable methylation profiling and want early access to next-generation epigenomic chemistry.
FAQs
1. What are the main benefits/advantages of using MeTLAND?
MeTLAND is a fast, user-friendly DNA methylation workflow that fits into any standard molecular biology lab.
It requires no harsh chemicals or specialized equipment and preserves DNA integrity throughout the process.
Key advantages:
> Requires only ~30M reads for high-quality data
> Rapid library preparation with minimal hands-on time
> Enables discovery-driven analysis with high-complexity libraries
2. What are the current offerings at NEXUS Epigenomics?
MeTLAND Early Access Program launched in Q4 2025 for selected researchers to benchmark our chemistry.
3. What types of samples can be used with MeTLAND?
MeTLAND works across a wide range of inputs including cell lines, primary cells, nuclei suspensions, fresh or frozen tissues. We are also optimizing compatibility with FFPE (formalin-fixed, paraffin-embedded) samples.
4. What is the minimum starting material needed for MeTLAND?
Cell suspensions: 200,000 – 250,000 cells.
Tissues: Varies with complexity, rigidity, and density; as a general guide, start with ~100-150 mg.
5. What is the cost?
Please contact us for current pricing.
We offer competitive pricing tailored to project size, sample volume, and workflow needs.